We’re excited to announce that BioLM now ships a production MCP (Model Context Protocol) server at https://biolm.ai/mcp and biolm/biolm-mcp on NPM. This means you can connect Claude, ChatGPT, and any MCP-compatible AI assistant directly to BioLM’s suite of in-silico modeling tools — no API glue code required.
What Is MCP?
Model Context Protocol is an open standard that lets AI assistants call external tools and APIs in a structured way. Instead of writing integration code, you point your AI client at an MCP server URL and it discovers the available tools automatically. The AI can then call those tools mid-conversation, reason over the results, and iterate, all without leaving the chat.
In protein engineering, that could look something like asking an agent to do the following:
1. Fetch a public PDB or sequence
2. Collect all known similar variants
3. Predict the scores of all variants using multiple models
4. Design new variants that exceed the scores of known variants, iterating until complete.
You can see an agent interacting with the BioLM MCP to do something like this below:
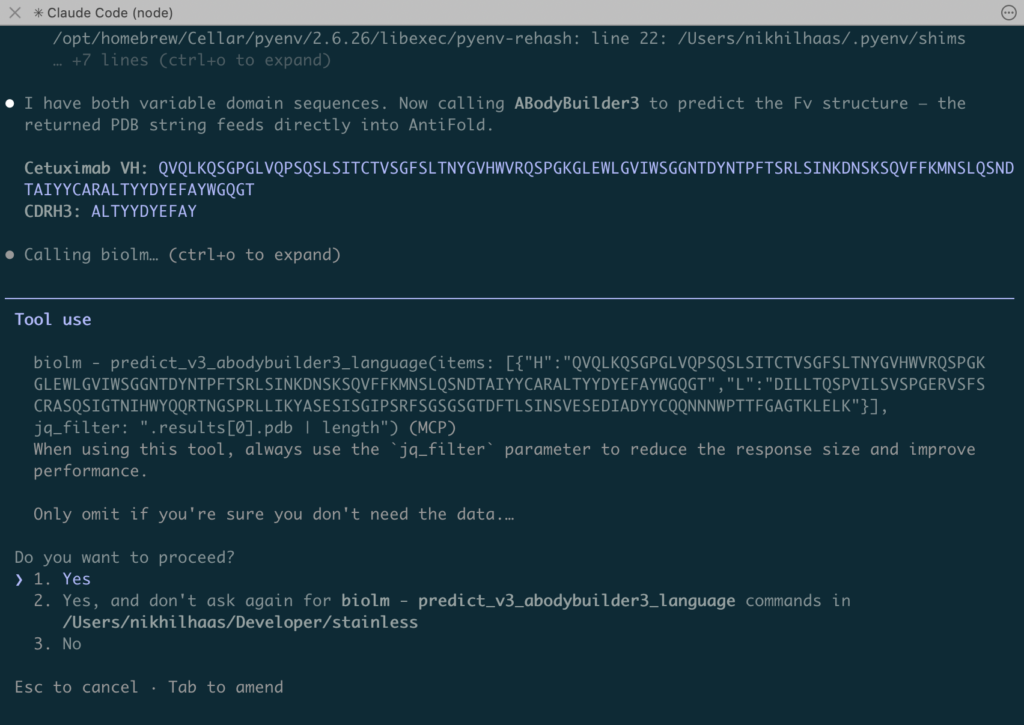
Why Use MCP
As large-provider foundation models like Sonnet, Opus, GPT 5, and GPT Codex have become more sophisticated at coordinating complex work. Compared to one year ago, model contexts up to 1M tokens have now become more common. This allows us to load the model with more info about how to find the right tool. In this case, that means finding the right model and understanding the tool’s request and response schema. This adds to the token used, but helps the model make a successful tool call and parse the response correctly.
What You Can Do with The MCP
Out of our 74+ models (Q2 2026), our initial launch of the BioLM MCP server exposes 20 tools covering the following workflows in computational biology:
| Category | Tools |
|---|---|
| Protein structure | ESMFold, ABodyBuilder3, NanobodyBuilder2 |
| Sequence embeddings | ESM-2 (150M), ESM-C (300M, 600M) |
| Variant effect | ESM-1v (ensemble) |
| Antibody annotation | SADIE, AbLang-2, IgBert (paired & unpaired) |
| Antibody design | AntiFold generate, ABodyBuilder3 |
| Peptide tools | Peptide encode |
| Nanobody tools | nanoBERT, NanobodyBuilder2 |
| Synthetic biology | DNA Chisel, Evo2 generate |
| Safety screening | BioLMTox-2 |
| Thermostability | TEMPRO 650M |
These are the same models that power BioLM’s API, now accessible as conversational tools.
What’s Different About BioLM’s MCP
While MCP sounds straightforward and standard in practice, utilizing a MCP server in a chat context requires MCP tool calls to respond quickly to meet the expectation of the user. This is a challenge for typical workflow-based platforms: tool calls to generate proteins or run predictions return an ID to a job that the agent has to poll for completion. In contrast, BioLM tool calls request real-time results from our GPU-backed APIs, in some cases returning results in a few seconds. This means less waiting for a result, more responsive agents, and less waiting for compute to complete.
As a by-product of BioLM models being served API-first, we have excellent request and response structure and validation for each model using Pydantic. You can see an example request and response schema and key descriptions on any of our model documentation pages, like these docs for Boltz-2. This provides excellent structure to create the BioLM MCP tools, and are enforced on each tool call. Pydantic provides informative responses indicating any issues with the request inputs so the agent can correct the input. This reduces errors and helps with proper interpretation of results by the agent.
Getting Started: Connect to The MCP
Quickly adding our MCP to your Claude or other coding agent config by following one of the options below.
Option 1: API Token
If you already have a BioLM API key, add the server to your AI client config with a single header:
{
"mcpServers": {
"biolm": {
"command": "npx",
"args": [ "-y", "@biolm/biolm-mcp", "--client=claude" ],
"env": {
"BIOLMAI_TOKEN": "YOUR_BIOLM_API_TOKEN"
}
}
}
}Replace YOUR_BIOLM_API_TOKEN with a token obtained from your BioLM account on your API Keys page.
Option 2: OAuth
BioLM’s MCP server supports full OAuth 2.0 with PKCE. When you add the server URL to a client like Claude or MCP Inspector, the client will redirect you to BioLM’s login page, obtain an access token, and handle token refresh automatically — no API key management needed.
For the local CLI, first run the command below to open a browser login flow and save credentials to ~/.biolmai/credentials . This provides authentication to the BioLM APIs for future sessions and tool calls:
npx @biolm/biolm-mcp --loginThen to register the server with Claude Code, the command would be:
claude mcp add biolm -- npx -y @biolm/biolm-mcp --client=claudeAdd -s user to make it global (all projects) rather than project-scoped:
claude mcp add -s user biolm -- npx -y @biolm/biolm-mcp --client=claudeThe –login step just writes credentials to ~/.biolm/credentials; claude mcp add is what actually registers the BioLM MCP in ~/.claude.json so Claude Code knows how to start its MCP server.
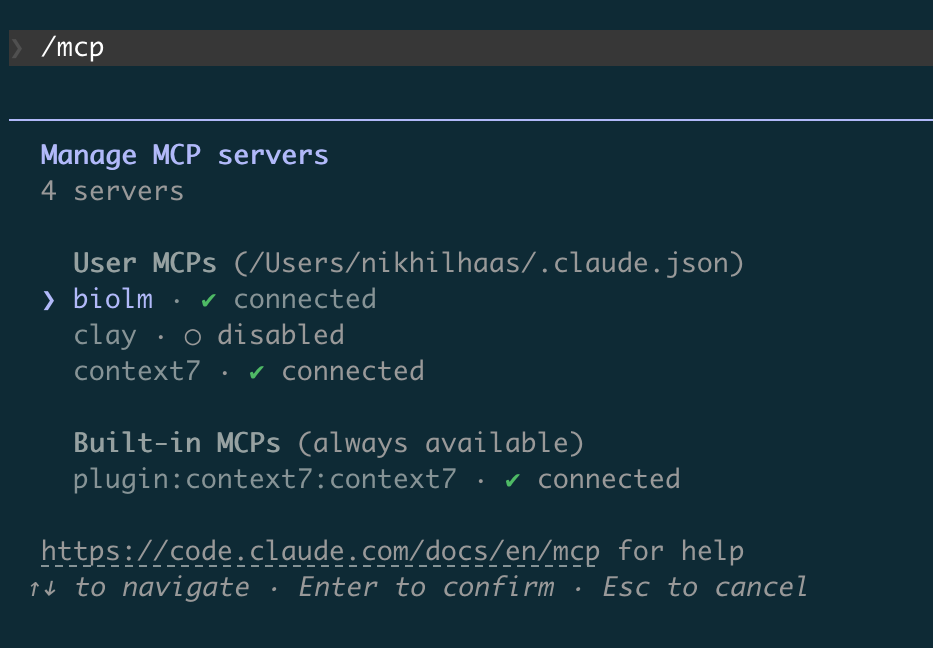
For both Options, in a Claude CLI chat, running /mcp should show the “biolm” MCP server connected:
Provider-Specific Configurations
Claude Desktop
On an Apple computer, add to ~/Library/Application Support/Claude/claude_desktop_config.json:
{
"mcpServers": {
"biolm": {
"command": "npx",
"args": [ "-y", "@biolm/biolm-mcp", "--client=claude" ],
"env": {
"BIOLMAI_TOKEN": "YOUR_BIOLM_API_TOKEN"
}
}
}
}Replace YOUR_BIOLM_API_TOKEN with a token obtained from your BioLM account on your API Keys page.
Filtering to Just the Tools Needed
The remote MCP URL supports query parameters for filtering tools by tag, resource, or name. This is useful for narrowing down what an agent sees. For example agents might make such queries to narrow down the available tools for a given task:
https://biolm.ai/mcp?tag=antibody
https://biolm.ai/mcp?resource=esmfold&resource=esm2_150m
https://biolm.ai/mcp?tool=predict_v3_esmfold&tool=predict_v3_esm1v_allOr it might query the search_docs tool in the BioLM MCP. Both methods are automatically available to your agent, and help it determine the best tools for the type of molecules you are working with.
Try It Now
- Visit biolm.ai to get an API key or create an account
- Add
https://biolm.ai/mcpto your MCP client - Ask your AI assistant: “Fold the protein sequence MENDELMENDEL with ESMFold and tell me the mean pLDDT”
- Contribute code for the BioLM MCP server on GitHub: BioLM/biolm-mcp
The BioLM MCP server is live and production-ready. This initial release primarily contains models with the shortest response times. We’re continuously expanding the tool set, supporting more of our models via MCP, and adding the ability to run larger-scale computational Protocols via MCP. If there’s a model or tool you’d like to see added, reach out to us at support@biolm.ai.